Study sites and bumblebee sampling
We conducted bumblebee sampling in both urban and rural areas across three Swiss regions (referred to as regions, Fig. 1), specifically within the Cantons of Basel, Bern, and Zurich54. For each region, three sampling sites were selected in urban areas (i.e., the cities of Basel, Bern, and Zurich) and in rural areas, except in rural Bern where only one site could be chosen (total = 16 sites, see section on Data records), following the methodology of Eggenberger et al.54. Specifically, urban sites were characterized by at least 60% impervious surfaces and were situated in the city centers, at least 1 km away from suburban areas and urban forests. Rural sites were standardized based on specific criteria: low-density settlements, meadow and pasture lands with similar management regimes, altitudes comparable to urban sites (400–600 m), proximity to water, and minimal forest cover. These rural sites were randomly selected within a defined area with a 4 km radius, representing the maximum extent of the urban centers. In each region, a minimum distance of 20 km separated urban and rural sites to minimize gene flow and interbreeding between bumblebee populations in these environments. The study encompassed 16 distinct sites: six around Zurich, six around Basel, and four around Bern. Within each region, we selected three non-overlapping sampling sites (radius of 800 m) that corresponded to the upper foraging ranges of the two target bumblebee species. Sampling began in the canton of Zurich, with the urban site in Zurich city (47°22′N, 8°33′E) and the rural site in the lower Töss valley (47°21′1′′N, 8°56′6′′E). The second region was in Basel, with the urban site in Basel-Stadt (47°34′N, 7°36′E) and the rural site in the Fricktal area in canton Aargau (47°30′21.43′′N, 8°3′7.63′′E). The third region was the canton of Bern, where the urban site was in the city of Bern (46°57′N, 7°27′E) and the rural site in the Bernese Mittelland (46°56′N, 7°12′E).
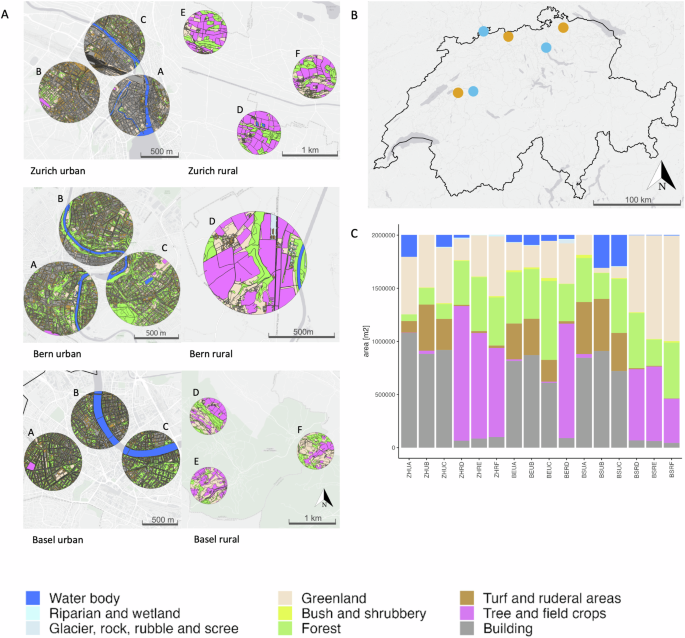
Sampling sites. (A) Configuration of the different habitats of the 16 sites across 6 regions sampled by Eggenberger et al.54 using the habitat map of Switzerland V1 (Price et al., 2021) in the cantons of Basel, Bern and Zurich. The 9 habitat tapes are indicated by colour. (B) Geographical distribution of the sampling regions in Switzerland (urban – blue, rural – orange). (C) Proportion of each habitat per site.
We focused on two bumblebee species, the common carder bee Bombus pascuorum (Scopoli, 1763) and the red-tailed bumblebee Bombus lapidarius (Linnaeus, 1761). Both species are widespread in the Swiss lowlands and occur in both urban and rural areas. While both are generalists, B. pascuorum has a longer tongue than B. lapidarius. Bumblebees were collected using hand-netting during the peak activity months for both species, from July to mid-August 2016. At each 800-m site, 30–40 individuals of each species were collected, with the exception of one urban site in Bern, where only three B. lapidarius individuals were found. Sampling efforts were standardized across all sites and conducted during peak bumblebee activity hours (09:00–17:00) under optimal weather conditions. We surveyed the entire 800-m radius for the target species but limited collection to a maximum of ten individuals per specific location within the site. Only active foragers were collected. Species identification of all collected individuals was confirmed in the laboratory, and specimens that could not be conclusively identified were excluded. Bumblebee samples were stored at -20 °C until pollen collection.
Pollen collection
We extracted pollen from the leg-baskets (i.e., corbicular pollen) and the body (i.e., body pollen) of the bumblebees (Fig. 2). For pollen collection, the bees were first removed from the -20 °C storage compartment.
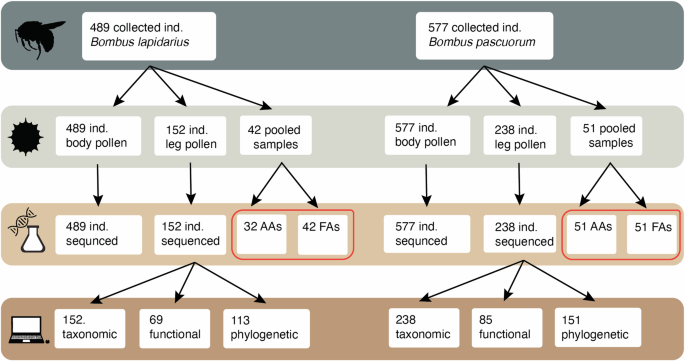
Summary of the workflow and gathered data for the two bumblebee species studied. The workflow is classified in four steps, from top to bottom: bumblebee individual collected, pollen extraction, metabarcoding and chemical analyses, and diet breadth calculations. For each step, the number of samples included is provided. From the starting number of individuals (489 individuals for Bombus pascuorum and 577 individuals in Bombus lapidarius) we used all individuals for the metabarcoding of pollen carried on the body of the bumblebees and a subset for the metabarcoding of the pollen carried in the leg-baskets (152 individuals in B. lapidarius, 238 individuals in B. pascuorum). Furthermore, we pooled the leg-basket pollen from different individuals from the same species collected in the same study site for the nutritional analyses (42 pooled samples for B. lapidarius and 51 pooled samples for B. pascuorum). From these samples, we uncovered the content and ratios of amino acids (AAs) and fatty acids (FAs), represented inside a red box in the figure. Finally, after obtaining the plant composition in the pollen collected in the leg-baskets, we estimated the diet breadth using taxonomic, functional and phylogenetic diversity metrics. Note that for B. lapidarius, we could only analyse the AAs in 32 out of 42 samples due to insufficient pollen mass.
Corbicular pollen
We extracted corbicular pollen in152 individuals of B. pascuorum and 238 individuals of B. lapidarius (Table 1, Fig. 2) across all sampling sites (Table 1, Fig. 2). To collect the corbicular pollen, the rear legs of each bumblebee were carefully detached under a binocular microscope, and the bee’s body was placed in a 2 ml Eppendorf tube. If pollen was present in the corbicula, it was gently removed using tweezers and a needle under the microscope and transferred to a prepared, well-labeled PCR plate. The storage vial was also checked for any corbicular pollen adhering to its sides (i.e., compact, moist pollen which could be clearly associated with the corbicular pollen), which was added to the leg samples. If no pollen was detected on the legs or in the vials, this step was omitted. We first extracted the corbicular pollen before extracting the body pollen.
Body pollen. We extracted body pollen on all 1066 bumblebee individuals (Table 1, Fig. 2). To collect the pollen, 500 μl of Milli-Q water was added to the bee in the Eppendorf tube, which was then briefly centrifuged and placed in an ultrasonic bath for 4 minutes. Following this, the Eppendorf tubes containing the bees were centrifuged at 10,000 RCF for 5 minutes. The bee was then removed and stored again at -20 °C. The remaining liquid and pollen mixture was centrifuged once more at 10,000 RCF for 1 minute to form a pellet, with the excess liquid discarded. The remaining liquid was then mixed with the pollen pellet and transferred to the PCR plate. The PCR plates were sealed with air-permeable tape and stored at -80 °C for at least 1 hour before being lyophilized overnight. The samples were then ready for metabarcoding and chemical analysis.
DNA metabarcoding & bioinformatics
DNA metabarcoding (isolation, amplification, and sequencing) was performed by AllGenetics laboratories (AllGenetics & Biology SL; A Coruña, Spain). DNA was isolated from samples using the Quick-DNA Microprep Plus Kit (Zymo Research), strictly following the manufacturer’s instructions. DNA was resuspended in a final volume of 15 μL. A DNA extraction blank (Bex1-16) in each round of the DNA isolation procedure was included, to be treated as regular samples in the next step of the library construction process to check for cross-contamination. The isolated DNA was quantified by fluorimetry with Qubit, using the High-Sensitivity dsDNA Assay (Thermo Fisher Scientific). There was quantifiable DNA from most samples, except from 93 samples. These 93 samples were too low for Qubit quantification detection with the High-Sensitivity dsDNA Assay kit, meaning that the DNA values were below 0.1 ng/μL. Therefore, library construction may be compromised.
For library preparation, a fragment of the ITS2 genomic region (of around 300 bp) was amplified using the primers ITS_S2F55 and ITS4R56, as proposed and previously applied to metabarcoding57. Illumina sequencing primers were attached to these primers at their 5’ ends. Then, PCRs were carried out in a final volume of 12.5 μL, containing 1.25 μL of template DNA, 0.5 μM of the primers, 6.25 μL of Supreme NZYTaq 2x Green Master Mix (NZYTech), and ultrapure water up to 12.5 μL. In the next step, the reaction mixture was incubated as follows: an initial denaturation step at 95 °C for 5 min, followed by 35 cycles of 95 °C for 30 s, 51 °C for 45 s, 72 °C for 30 s, and a final extension step at 72 °C for 10 min. The oligonucleotide indices which are required for multiplexing different libraries in the same sequencing pool were attached in a second PCR round with identical conditions but only 5 cycles and 60 °C as the annealing temperature. A negative control that contained no DNA (BPCR) was included in every PCR round to check for contamination during library preparation. The libraries were then run on 2% agarose gels stained with GreenSafe (NZYTech), and imaged under UV light to verify the library size.
The libraries were purified using the Mag-Bind RXNPure Plus magnetic beads (Omega Biotek), following the instructions provided by the manufacturer. Then, the libraries were pooled in equimolar amounts according to the quantification data provided by the Qubit dsDNA HS Assay (Thermo Fisher Scientific). This pool also contained a testimonial amount of the corresponding extraction blanks (Bex) and the PCR blanks (BPCR). The pool was sequenced in a fraction of an Illumina NovaSeq PE250 flowcell (Illumina), aiming for a total output of 30 gigabases.
For bioinformatics, we followed the pipeline outlined at https://github.com/chiras/metabarcoding_pipeline58. This pipeline was implemented using VSEARCH v2.14.259 for merging, quality truncation, and filtering (maxEE < 1; sequence length between 150 bp and 300 bp). Cleaned reads were then denoised into amplicon sequence variants (ASVs) and filtered for chimeras using VSEARCH. The ASVs were initially mapped using global alignments with VSEARCH against a floral ITS2 reference database specific to the study region, with an identity threshold of 97%. This database was created using BCdatabaser60 and included a list of potential plants found in the study area. For reads that remained unclassified, we used SINTAX61 to assign the deepest possible taxonomic levels using a global reference database62.
In DNA metabarcoding studies it has been observed that a low percentage of the reads of a library can be assigned to another library. This phenomenon, referred to as mistagging, tag jumping, index hopping, index jumping, etc. is the result of the misassignment of the indices during library preparation, sequencing, and/or demultiplexing steps63,64,65,66. ASVs were aggregated to species or genus level, depending on the deepest classification. In order to correct for this phenomenon, species occurring at a frequency below 0.01% in each sample were removed.
Nutritional analyses
We focused on two critical macronutrients for bumblebee health and fitness: amino acids and fatty acids67. In order to have a sufficient pollen mass to perform the nutritional analyses, we pooled pollen (from the leg-baskets) from different bumblebee individuals collected by Eggenberger et al.54 within study sites and for each species separately. Overall, this resulted into 92 samples for fatty acid analyses and 85 for the amino acid analyses (Fig. 2).
Solvents and reagents needed for nutrient analyses
For the amino acid analysis:
For the fatty acid analysis:
Amino acid analysis
We conducted amino acid analysis using ion exchange chromatography (IEC) with a Biochrom 20 plus amino acid analyzer, following the protocol by Kriesell et al.68. To analyze protein-bound amino acids (AA) in pollen, 5–10 mg of the collected pollen was first extracted in an ultrasonic bath with 100 µl of deionized water for 30 minutes. The extract was then refrigerated for 60 minutes, followed by centrifugation and membrane filtration for 10 minutes. The remaining residue, which was reserved for protein-bound amino acid analysis, was treated with 200 µl of 6 N HCl, boiled at 100 °C for 4 hours, cooled to room temperature, and centrifuged for 10 minutes. The supernatant was then subjected to evaporation at 100 °C to remove water and HCl, followed by re-dissolution and boiling in 200 µl of fresh water until dry. This process was repeated twice to ensure complete removal of the acid, but during the last time the sample was then centrifuged again before water was evaporated.
Next, 100 µl of the supernatant was mixed with 20 µl of 12.5% sulfosalicylic acid, frozen overnight, and extracted in the refrigerator for 30 minutes the following day. The mixture was briefly stirred, centrifuged for 10 minutes, and 100 µl of the supernatant was combined with 100 µl of sample rarefaction buffer (lithium buffer). This solution was then membrane-filtered by centrifugation for 5 minutes, and 20 µl of the filtrate was diluted in 80 µl of sample rarefaction buffer for IEC analysis. To quantify amino acids, we used the area of the peaks, calibrated via an external standard (physiological calibration standard from Laborservice Onken GmbH, Gründau, Germany) containing all proteinogenic amino acids except glutamine and asparagine, which were manually added before running the standards and samples (standard concentration was 200 µM per amino acid). The column used was a PEEK column packed with cation exchange resin. Amino acids were derivatized post-column via ninhdydrin and recorded via UV detection. Note that tryptophan could not be analyzed because it is destroyed by HCl.
Fatty acid analysis
Fatty acid (FA) analysis followed the protocol by Villagómez et al.69. For each pollen sample, 0.5 mg was homogenized with 7 µl of a 200 ng/µl solution of nonadecanoic acid in chloroform (used as an internal standard) in 0.1 ml of a 2:1 mixture of chloroform and methanol (both from Sigma-Aldrich, Taufkirchen, Germany). An additional 0.4 ml of the chloroform-methanol mixture was then added for further homogenization, and the mixture was transferred to a new vial with another 2.5 ml of the chloroform-methanol mixture, resulting in a total volume of 3 ml. The samples were shaken at 250 rpm for 24 hours and then evaporated to dryness. Subsequently, 10 µl of trimethylsulfohydroxide (TMSH) in 150 µl of dichloromethane (both from Sigma-Aldrich) was added, and the samples were analyzed using a gas chromatograph (GC, Agilent Series 8890) coupled to both a mass spectrometer (MS, Agilent 5977 C) and a flame-ionization detector (FID). Helium was used as the carrier gas (1 mL/min), and 1 µl of the sample was injected in splitless mode at 320 °C onto a fused silica column (Agilent Technologies DB5, 30 m length, 0.25 mm diameter). The oven temperature was initially set at 60 °C, increased to 150 °C at a rate of 15 °C/min, held for 10 minutes, then increased to 320 °C at 10 °C/min, and held for another 10 minutes.
Fatty acids were identified by comparing the mass spectra and retention times of peaks in the resulting chromatograms (MS) to standards (e.g., FAME C8-C24 and single fatty acid standards, Sigma-Aldrich), while the FID chromatograms were used to quantify fatty acids using the internal standard. In this method, di- and triglycerides are broken down into fatty acid methyl esters, so the FA content in this study refers to the total content of both free and glyceride-bound FAs. We also calculated the content and ratios of specific types of FAs relevant to bee nutrition, survival, and health70.
Metrics on nutrient intake
The total protein content was calculated as the sum of all amino acids, and the amino acid (AA) content in this study refers to the total protein-bound AA content. Additionally, we calculated the total content of essential AAs, which animals cannot synthesize and must obtain from their diet, as well as non-essential AAs, which animals can synthesize. The ratio between essential and non-essential AAs was also calculated. We calculated the content of saturated, unsaturated, omega-3, omega-6, and omega-9 FAs, as well as the ratios of saturated to unsaturated FAs and omega-3 to omega-6 FAs. These FA metrics are significant indicators of nutrition and health.
Metrics on diet breadth
We utilized taxonomic, functional, and phylogenetic metrics to assess bumblebee diet breadth at individual level (Table 2). Plant species richness served as the metric for plant taxonomic diversity71. For functional diversity, we calculated multidimensional metrics, focusing on three key dimensions: functional richness, functional evenness, and functional divergence, using the “FD” package (version 1.0–12.1) by Laliberté & Legendre72. We extracted plant traits (i.e., flowering duration18, flowering start18, growth form18, structural blossom class18, sugar concentration51,52, symmetry18 and plant height18) from multiple published, open-source datasets18,51,52. To compute functional diversity indices, we included flowering duration, structural blossom class, and pollen sugar concentration, as other traits present in the datasets (i.e., symmetry, flowering start, and plant height) had high correlations ( > 0.7). Moreover, besides the correlations, we limited the number of traits to three because the number of plant species in the pollen needed to exceed the number of traits, and most bumblebee individuals carried fewer than four plant species. Consequently, growth form was excluded to avoid filtering out too many bumblebee individuals.
For phylogenetic metrics, we calculated multidimensional indices, specifically phylogenetic variability, phylogenetic richness, phylogenetic evenness, and phylogenetic clustering, using the “picante” package (version 1.8.2) by Kembel et al.73, with the phylogeny from Jin and Qian71. Bumblebee individuals with fewer than four plant species in their collected pollen were excluded from the functional diversity analysis, as the convex hull could not be computed. For phylogenetic diversity metrics, individuals with fewer than three species were removed. In total, this resulted in 154 bumblebees being included for functional diversity analysis and 264 individuals for phylogenetic diversity analysis (Fig. 2).
link











More Stories
Sugary drinks linked to rising anxiety in teens
Planetary Health Diet Study Highlights Nutrient Tradeoffs
What happens to the body on the carnivore diet?